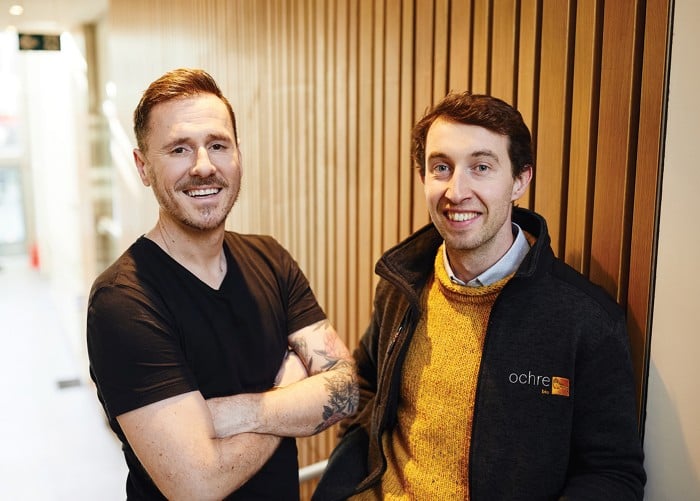


About Ochre Bio
The Ochre Bio team comprises a diverse, interdisciplinary group of scientific experts working on chronic liver diseases. They faced a challenge in moving some of their work forward because of a dependency on a manual image-processing workflow, compounded by working across different locations and time zones.
The Challenge
One of Ochre Bio’s Taipei-based computational biologists would run a workflow to support a team of wet lab scientists based in New York City (NYC). This involved image pre-processing in Python to divide large images into smaller ones, characterizing image tiles using the open-source CellProfiler software, then aggregating results and generating plots for visualization with R.
Created separate Compute Capsules for image preprocessing, running CellProfiler, and results visualization
Created a Pipeline to run multiple instances of the CellProfiler Capsule in parallel on AWS Batch

Created a parameterized no-code version of the pipeline to share with NYC-based colleagues who could use it with no additional support
Ochre Bio automated their manual workflow by moving it into Code Ocean
The Results
“Code Ocean sped up our internal image pre-processing computational workflow by at least 10x, improving collaboration and productivity between our global teams across time zones and removing the need for complicated and painful cloud computing infrastructure set up.”

Dimitris Polychronopoulos
Director of In Silico Biology, Ochre Bio
Key features used
Key integrations








 Capsules
Capsules
 Pipelines
Pipelines
 Data assets
Data assets
